PacBio Newsletter_Feb 2021
2021-02-09 09:45:04 Source:Gene Company Limited



PacBio announced a multi-year collaboration with Invitae Corporation, a leading medical genetics company, to begin development of a production-scale high-throughput sequencing platform leveraging the power of PacBio’s highly accurate HiFi sequencing to expand Invitae’s whole genome testing capabilities.
With the ability of HiFi reads to detect genetic variants and even in hard-to-sequence regions of the genome, this collaboration make WGS-powered analysis more accessible for areas such as carrier screening, evaluating immune system responses, and diagnosing other heritable disease.
For more details, please click HERE.


PacBio APAC Virtual HiFi Sequencing Summit 2020
Please click Here to watch. If you have not registered yet, please register according to the website’s instruction prior to access the webinar.

PAGBio Day 2021 (PacBio Plant and Animal Genomics Day)
Please click Here to watch. If you have not registered yet, please register according to the website’s instruction to access the webinar

HiFi Metagenomics 101: From Sample to High Resolution Genomes
Please click Here to watch.
Protocol/document Updates
Protocol/document releases from Dec 2020.
Protocols | Released date | Changes |
January 26, 2021
| Initial release | |
Procedure & Checklist – Amplification of Full-Length 16S Gene with Barcoded Primers for Multiplexed SMRTbell Library Preparation and Sequencing | January 7, 2021 | Added 12 more Reverse Primers for a total of 24, allowing multiplexing of up to 192 samples. |
Procedure & Checklist – Preparing SMRTbell Libraries using PacBio Barcoded M13 Primers for Multiplex SMRT Sequencing | December 18, 2020
| Initial release. |
Procedure & Checklist – No-Amp Targeted Sequencing Utilizing the CRISPR-Cas9 System | December 15, 2020 | Updated to support 48 plex. Provided additional Guide RNAs and Barcoded Adapters. Procedure is updated to improve flow. |
Overview – Sequel Systems Application Options and Sequencing Recommendations | December 3, 2020 | Added information including IPA for genome assembly, pbaa for amplicon analysis, no amp workflow is now supporting up to 48 samples/ cell |
Publication Highlights

PNAS February 2, 2021 118 (5) e2019768118. https://www.pnas.org/content/pnas/118/5/e2019768118.full.pdf

•Chinese University of Hong Kong (CUHK) research team led by Prof Dennis Lo have developed a new method that greatly improves the detection of DNA cytosine methylation using Pacific Biosciences single-molecule real-time sequencing data.
•The approach, called the holistic kinetic (HK) model considers kinetic signals from the DNA polymerase used in PacBio sequencing, as well as the sequence context, to determine cytosine methylation sites.
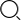
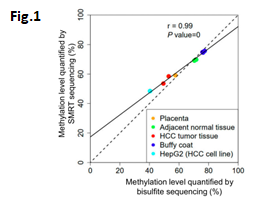
•After training a methylation classification model using a convolutional neural network, it was reported being able to detect cytosine methylation, or 5-methyl-C, with 90% specificity and 94% sensitivity, with a 99% correlation of overall methylation level with bisulfite sequencing (Fig.1).
•The advantage of the method is that it allows simultaneous determination of DNA sequence and CpG methylation in one go, with high base-calling accuracy, and without the use of a step that is destructive to the input DNA which is always a problem with the traditional method of bisulfite sequencing.
•More importantly, PacBio long read sequencing platform can overcome the problem of short read sequencing platform on generating methylation data over larger distances in the genome to generate methylation haplotypes.
•The potential applications of the patent-pending method include epigenetic studies in humans and other organisms as well as molecular diagnostics.