PacBio Newsletter (Issue July)
2020-07-14 08:26:35 Source:Gene Company Limited
Monthly Newsletter Jul, 2020
Pacbio Updates
NEW! SMRT Sequencing Datasets Webpage
All Sequel II generated publicly available datasets are released on PacBio webpage. This includes applications in Whole Genome Sequencing, RNA sequencing, Targeted Sequencing and Complex Populations.
Please click here to explore.

COVID-19 Update
New Protocol for Sequencing SARS-CoV-2 without PCR Amplification is released.
Cultured virus protocol: VAST/Pasteur Method for SMRT-Sequencing of SARS-CoV-2 from Cultured Virus Supernatant.
2020 Plant & Animal Sciences SMRT Grant Program
To win the library construction, HIFI sequencing on the Sequel ® II System, and bioinformatics analysis, please submit your 90-second video abstract to this website by July 24, 2020!
Find out more in this video here.
PAY attention! SMRT Link V9.0 is Released
Improvements
• CCS Analysis is 15-20% faster!
• Enhanced Analysis applications – Microbial Assembly and Base Modification analysis, and support for the new Ultra-low DNA input protocol
• GUI support for up to 10,000 multiplexed samples.
Please refer to Installation Guide here to upgrade from SMRT Link V8 or for a fresh install.
Protocol/document Updates
Protocol/document releases from Jun 2020.
Protocols | Released date | Changes |
Procedure & Checklist – Preparing SMRTbellLibraries using PacBio Barcoded Overhang Adapters for Multiplexing Amplicons | June 17, 2020 | Updated required materials table on page 2.
|
Procedure & Checklist – Preparing HiFi Libraries from Low DNA Input Using SMRTbell Express Template Prep Kit 2.0 | June 30, 2020 | •Low DNA Input protocol has been updated and changed to CCS application instead CLR, thus HiFi reads will be used for further analysis. Evaluation of gDNA sample has been updated to meet the requirements for HiFi library construction (Page 6-7). |
NEW! DNA Resource
A new Technical Note about DNA extraction and QC for HIFI sequencing is released
Click here to find this document. 
Publication Highlight

Liu et al., 2020, Cell. 182, 1–15.
https://doi.org/10.1016/j.cell.2020.05.023

lDe novo genome assemblies for 26 representative wild and cultivated soybeans were performed by PacBio SMRT sequencing with the aid of optical mapping by Bionano Irys System
lA graph-based pangenome is constructed which identified numerous genetic variations that cannot be detected by short read sequencing
lIdentification of thousands of structural variants and dozens of gene fusions
lLinked the structural variations to gene expressions and agronomic traits
lThis pan-genome resource help to promote evolutionary and functional genomics studies in soybean
New webinars
No Organism Too Small – Build high-quality genome assemblies of small organisms with HiFi sequencing (72mins)
- By Scott Geib, Christopher Laumer, and Marcela Uliano da Silva and Michelle Vierra
•Overcome the challenges of sequencing organisms with small body size using the new low and ultra-low DNA input methods from PacBio
•The advantages of using HiFi reads to sequence and de novo assemble genomes of single individuals

Click here to watch.
Increasing solve rates for rare and Mendelian diseases with long-read sequencing (68mins)
-By Kristen Sund and Aaron Wenger
•Recommendations on how to design your own study using HiFi reads
•Use long-read sequencing to solve rare neurological diseases involving complex structural rearrangements that were previously unsolved with standard methods.

Click here to watch.

Automated Capillary Electrophoresis
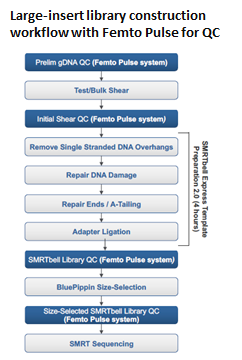
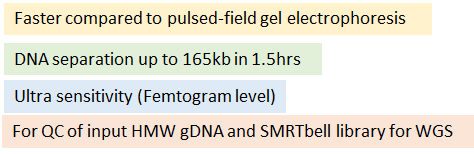
Highly recommended items for Quality Control (DNA sizing and quantification) in library preparation for SMRT sequencing
Femto Pulse



Fragment Analyzer






